Before we begin, familiarize yourself with these acronyms for a more pleasurable reading experience:
- RAG: Retrieval Augmented Generation
- API: Application Programming Interface
- BERT: Bidirectional Encoder Representations from Transformers
- GPT: Generative Pre-trained Transformer
Hello loyal readers!
(*crickets chirp*)
I’ve been working on a little personal project, a (small) medical coder, in order to familiarize myself with the practical implementation of a few concepts I had been reading about in theory, specifically, LLMs and retrieval-augmented generation. I also used this project as an excuse to test out some new tools I had been wanting to try (Carbontracker and ThauraAI). Given that my main domain of interest is healthcare, I also wanted to keep moving and learning in that direction.
What does it do?
Doctors write notes like: “Patient has chest pain, diagnosed with heart attack” which need to be converted to ICD-10 codes. This is called medical coding and is usually done by humans, but humans can make mistakes, take a lot of time, and cost a lot of money. Wouldn’t it be nice to have some supplemental help with this task? This model will automatically suggest correct ICD-10 codes based on doctor notes.
Here, we will predict the primary diagnosis only, so one code per clinical note (a single-code predictor, if you will) whereas, in reality, usually clinical notes include many codes.
Some notes
First, I want to note that I initially implemented the OpenAI API GPT-3.5 Turbo model (with a $5 limit) out of convenience, but soon after decided to try other options because, although it worked, I wanted to be more intentional about the tech companies that I use and support. And so, I switched to ThauraAI.
ThauraAI is an AI chatbot created as an ethical alternative to ChatGPT by two Syrian engineers. It is significantly more environmentally sustainable than ChatGPT, takes data privacy seriously by never utilizing user data to train models, and is genuinely people- and planet-oriented. I highly recommend it, I’m genuinely a huge admirer of what they are doing, and I’m glad I was able to use it for this project.
Another tool I implemented was Carbontracker, which tracks carbon intensity and energy consumption during model training. It’s very user friendly and I’ve been wanting to try it ever since it was mentioned at a FAIMI virtual workshop I attended. (Note that it only tracks the energy consumption and carbon footprint of my local CPU and not that of the ThauraAI API.)
How does it work?
RAG is an AI framework that works with LLMs to improve accuracy of responses. It Retrieves relevant information, Augments the model’s knowledge with fresh info, and Generates improved responses. Essentially, it can give a pretrained LLM more domain-specific knowledge so it can output more accurate info and “hallucinate” less, all without any retraining.
First, the knowledge base is created from which we will be retrieving the aforementioned relevant information from. In this case, it is composed of synthetically generated clinical documents and of ICD-10 guidelines accessed via the MIMIC-IV database. These guidelines are definitions of diseases with their corresponding codes. We generate synthetic clinical notes through the ThauraAI API, which allows me to send text over the internet and get a response back from ThauraAI instead of running a huge LLM on my little baby laptop.
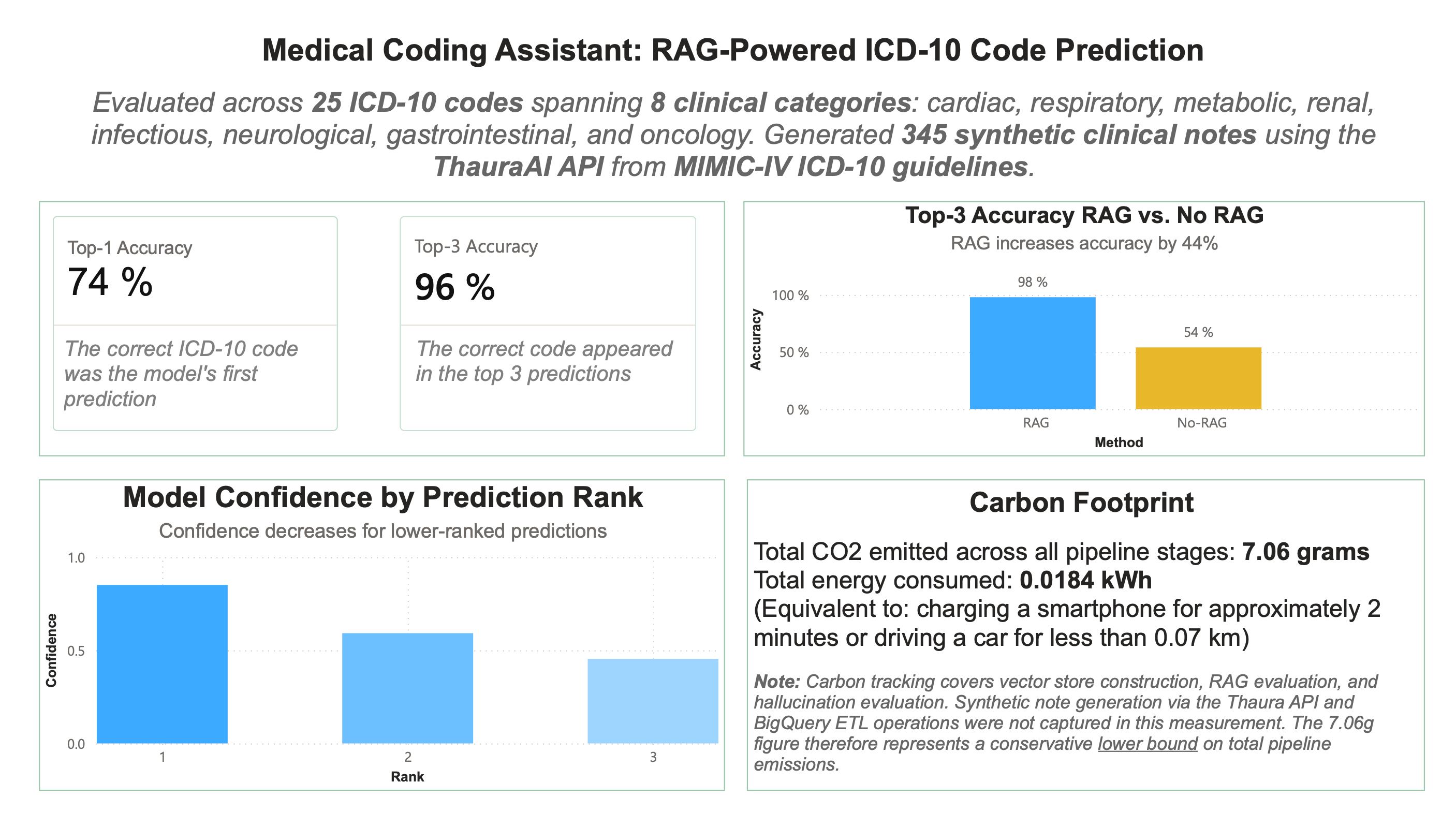
We use synthetic notes instead of real clinical notes from MIMIC-IV which, though de-identified, could still be in violation of the Physionet data-usage agreement if sent to a third-party application, in this case ThauraAI. We reference 25 ICD-10 codes, spanning 8 clinical categories (cardiac, respiratory, metabolic, renal, infectious, neurological, gastrointestinal, and oncology) and for each of those codes, 15 clinical notes are generated describing a patient with that diagnosis, for a grand total of 375 synthetic clinical notes. We also specifically prompt the model to exclude the name of the disease in the clinical note and also run the note through a cleaning function before prediction time. This is to ensure that the LLM can’t just read the exact diagnosis in the note and must actually predict it. Then, that set of synthetic clinical notes is split up into a training set that goes into the knowledge base and a validation and test set which are used to evaluate the model on. After the model is given a detailed prompt including a synthetic clinical note from the validation set and relevant context that is retrieved from the knowledge base, it returns its top 3 ICD-10 code predictions.
The way that RAG is able to decide what context from the knowledge base is relevant is by calculating vector similarity between embeddings. Here, I implemented Bio-ClinicalBERT, which is a version of BERT trained on medical text, specifically clinical notes. It converts medical text into numbers (vectors) the computer can understand, aka embeddings, which are stored in ChromaDB. ChromaDB compares vectors based on L2 (Euclidean) distance or cosine similarity, not sure which actually, but the point is that it compares vectors based on how similar they are and outputs the top 20 most similar guidelines and notes. Then, cross encoder reranking is able to read documents as pairs and identify more nuanced similarities, so for each of the top 20 similar documents found by ChromaDB, the cross encoder model (ms-marco-MiniLM-L-6-v2) analyzes them and returns the top 10 most similar ones to the input note. The LLM sees the current clinical note and retrieved documents (10 docs = similar past cases + official ICD-10 guidelines), supporting the model in making a more accurate ICD-10 code prediction .
How did it do?
To evaluate the RAG implementation, I compared Top-1 Accuracy vs. Top-3 Accuracy. RAG outputs the top 3 codes it thinks that the note is indicating. (Is the correct code the top one? Is the correct code within the top 3?) I also analyze these results to see where the model is getting confused. Sometimes, errors are in predictions where the model believes the note to be indicating a disease that has similar symptoms to the actual diagnosis. Here, it reached 76% top-1 accuracy and 96% top-3 accuracy.
I also compared RAG to the LLM on its own to check for accuracy and hallucination improvement, hoping that RAG would improve both. Ultimately, RAG increased accuracy by 44% (impressive) and produced 2 fewer hallucinated codes than the LLM on its own (not that impressive, but better than nothing). However, this is to be expected, since RAG was limited to only predicting codes that were within the 25 included, so hallucinating a code that didn’t exist was impossible, meanwhile the LLM on its own did not have that constraint. So, upon reflection, I don’t think it was a great way to measure hallucinations as it wasn’t a fair comparison between the two since they had different constraints.
Some struggles
To generate the synthetic notes, I altered the prompt several times to improve the quality of the note because I was having some issues with note accuracy. For example, the note was meant to be for sepsis and, when I was still using OpenAI’s GPT, it was generating a note describing septic arthritis. However, this is an issue even humans face when differentiating between conditions that have similar symptoms.
Also, when I switched to ThauraAI, because it is a reasoning model, there was a lot of extra noise to sort through in order to get to the actual code prediction or the actual synthetic note, meaning I needed significantly more tokens to get it to work for me (20 for OpenAI vs. 2000 for ThauraAI).
Future expansion: Multi-code Prediction
It would be interesting to see how expanding to a multi-code prediction would go. Synthetic notes would be generated to describe three conditions and the model would predict the primary condition as well as two secondary conditions. The model would be evaluated by its accuracy in predicting primary code in addition to its accuracy in predicting the two secondary codes, with similar evaluation and hallucination checks as for the single-code predictor.
Additionally, this current medical coder is very small, some might even say miniscule, given that there are around 70,000 existing ICD-10 codes and here we took only 25 into account. It would definitely be illuminating to expand, but I don't know if my CPU could take a larger knowledge base or if my wallet could take buying any more tokens.
Conclusion
Even though I didn’t reach any show-stopping, groundbreaking results, I learned a few things and I had fun. I also have been wanting to play around in Power BI so I made this visualization with my results. In the future, I’ll hopefully be making some more interactive ones, but since this project was quite small, I think this simple summary of results is sufficient.
Woohoo. See you next post.
- https://arxiv.org/html/2005.11401v4
About RAG:


